Hickson Group
Our laboratory investigates the biochemical, molecular and cell biological functions of DNA repair factors in eukaryotic cells and how these factors act to maintain chromosome stability.
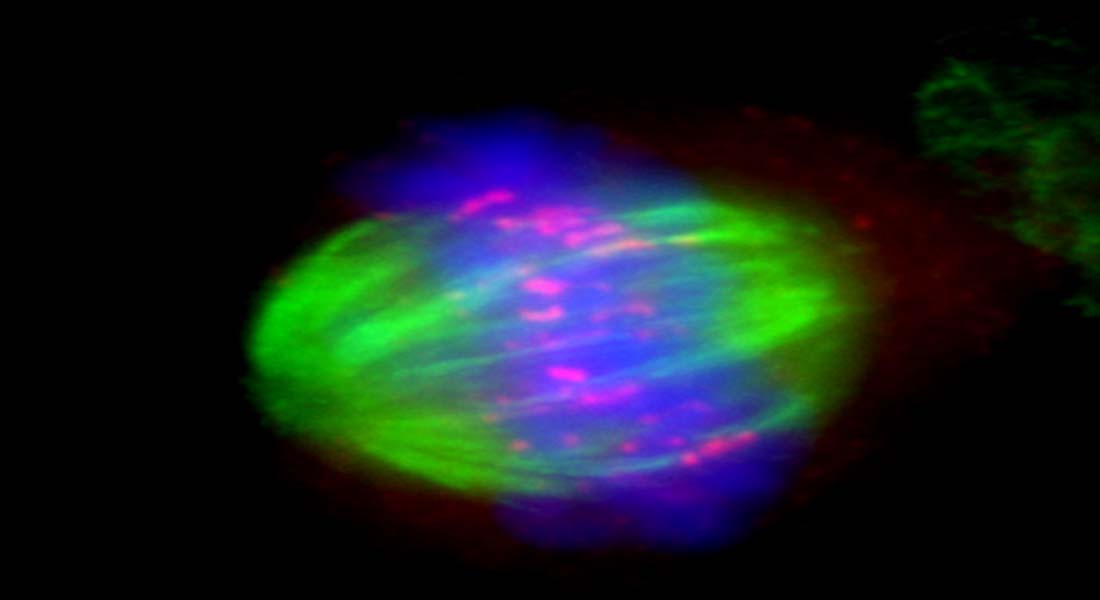
"Mechanism of chromosomal fragile site instability"
Our research is focused on understanding how chromosomal instability impacts on human disease. We study genes in human cells that, when mutated, lead to defective genome maintenance and high rates of cancer, neurodegeneration or infertility.
In recent years we have focused on common fragile sites as a model for understanding how ‘difficult-to-replicate’ regions of the human genome affect propagation of the genome through successive cell divisions. As part of these studies, we identified ultra-fine anaphase bridges (UFBs) which are DNA thread-like structures connecting the separating sister chromatids in the anaphase of mitosis. These UFBs exist in every human cell division but are normally ‘invisible’ due to their lack of staining with DNA dyes. They are coated with specific proteins, including the SNF2 family protein, PICH and the BLM helicase (which when mutated causes Bloom’s syndrome – a cancer predisposition disorder), that allows detection using specific antibodies.
We combine protein biochemistry with molecular/cell biology and high resolution imaging of human cells. Latterly, we have also been developing tools to exploit new developments in microfluidics and single molecule biophysical techniques to reconstitute in vitro defined steps in DNA metabolism.
Understanding how human cells complete DNA replication.
In 2015, our group identified a process that we named MiDAS (Mitotic DNA Synthesis) that serves to complete the duplication of certain regions of the human genome in the early stages of mitosis. MiDAS is a DNA repair-based process that takes over the task of completing the replication program at so-called ‘difficult-to-replicate’ loci such as common fragile sites (CFSs) and telomeres. CFSs are a major interest of the group as they are inherently unstable and are frequent sources of genomic alterations that arise in human cancers. We use molecular/cell biological techniques, coupled with genome-wide sequencing tools, to define mechanistically how MiDAS is conducted. Our current model is that MiDAS is a form of break-induced DNA replication (BIR) that requires homologous recombination factors and specialized DNA endonucleases and polymerases.
Defining the role of the PICH translocase and the Bloom’s syndrome complex in resolving ultra-fine DNA bridges in mitosis.
Ultra-fine anaphase DNA bridges (UFBs) were discovered in our group over 10 years ago. They are nucleosome-free DNA threads that connect the separating sister DNA masses during the mitotic anaphase. They arise from specific loci, including centromeres, CFSs and telomeres. UFBs cannot be detected using DNA dyes and are visualized using antibodies to bound proteins, including PICH and the Bloom’s complex (BLM, TOPOIIIa, RMI1, RMI2; BTRR). We are investigating why UFBs arise and how PICH and the BTRR complex acts to resolve them alongside TOPOIIa. We utilize cell biological techniques coupled with in vitro biochemistry using optical tweezers. This single molecule biophysical technology allows us to model the binding and resolution of entangled DNA molecules in real-time. In parallel, we study the cells of patients lacking functional BLM/TOPOIIIa, or laboratory-generated mutant mice or cell lines.
A novel system to manipulate and visualize human chromosomes.
In collaboration with physicists in Amsterdam (Gijs Wuite and Erwin Peterman) we are establishing methods to purify, grasp and manipulate intact human metaphase chromosomes using optical tweezers. With this, we aim to reconstitute sister chromatid disjunction and define how manipulating the degree of condensation of mitotic chromosomes affects their bio-mechanical properties.
Defining how DNA topology is regulated in human cells
Genomes are often compared to books, in which the information is encoded over several pages of A, T, G and C, readily accessible upon “keyword” request. In reality, genetic information is carried by an extraordinarily long, double helical DNA polymer that is highly compacted to fit into the tiny space of a cell nucleus. This inevitably leads to extensive DNA entanglement. Our cellular machineries, therefore, have to exploit, protect and copy topologically-entangled genomes. The interplay between these entanglements and DNA metabolism is largely under investigated, in spite of the fact that it represents a fundamental aspect of basic molecular biology. Indeed, little is known about the actual topology of the human genome, how it is regulated, and how cellular machineries exploit it to modulate processes such as transcription, repair, replication or chromosome architecture. Our aim is to generate new discovery tools to help increase our understanding of general supercoiling homeostasis in human cells and how it contributes to genomic stability maintenance. In particular, we combine biochemistry, biophysics and in vivo approaches to 1) identify of new actors of DNA topology, and 2) develop innovative strategies to monitor DNA topology in vivo.
- Rahul Bhowmick, Mads Lerdrup, Sampath Amitash Gadi, Giacomo G. Rossetti, Manika I. Singh, Ying Liu, Thanos D. Halazonetis, Ian D. Hickson. RAD51 protects human cells from transcription-replication conflicts. Molecular Cell, 2022, 2022 Aug 18;S1097-2765(22)00706-7
- Meijering, A.E.C., Sarlos, K., Nielsen, C.F., Witt, H., Harju, J., Kerklingh, E., Haasnoot, G.H., Bizard, A.H., Heller, I., Broedersz, C.P., Liu, Y., Peterman, E.J.G., Hickson, I.D.* and Wuite, G.J.L.* (2022) Nonlinear mechanics of human mitotic chromosomes. Nature 605, 545-550. *Co-corresponding authors
- Macheret, M., Bhowmick, R., Sobkowiak, K., Padayachy, L., Mailler, J., Hickson, I.D.* and Halazonetis, T.D.* (2020) High resolution mapping of mitotic DNA synthesis regions and common fragile sites in the human genome through direct sequencing. Cell Research, 30, 997-1008, *Co-corresponding authors.
- Xing, M., Zhang, F., Liao, H., Chen, S., Che, L., Wang, X., Bao, Z., Ji, F., Chen, G., Zhang, H., Li, W., Chen, Z., Liu, Y., Hickson, I.D.*, Shen, H.* and Ying, S.* (2020) Replication stress induces ATR/CHK1-dependent non-random segregation of damaged chromosomes. Molecular Cell, 78, 714-724. *Co-corresponding authors.
- Wu, W., Bhowmick, R., Özer, Ö., Vogel, I., Thakur, R.S., Richter, P.H., Ren, L., Hickson, I.D.* and Liu, Y.* (2020) RTEL1 resolves conflicts between DNA replication and transcription. Nature Struct. Molec. Biol., 27, 424-437. *Co-corresponding authors
- Bizard, A.H., Allemand, J-F., Hassenkam, T., Paramasivam, M., Sarlós, K., Singh, M.I. and Hickson, I.D. (2019) PICH and TOP3A cooperate to induce positive DNA supercoiling. Nature Struct. Molec. Biol., 26, 267-274.
- Sarlós, K., Biebricher, A.S., Bizard, A.H., Bakx, J.A.M., Ferrete-Bonastre, A.G., Modesti, M., Paramasivam, M., Yao, Q., Peterman, E.J.G., Wuite, G.J.L. and Hickson, I.D. (2018) Reconstitution of anaphase DNA bridge recognition and disjunction in vitro. Nature Struct. Molec. Biol., 25, 868-876.
- Bhowmick, R., Minocherhomji, S. and Hickson, I.D. (2016) RAD52 facilitates mitotic DNA synthesis following replication stress. Molecular Cell, 64, 1117-1126.
- Minocherhomji, S., Ying, S., Bjerregaard, V.A., Bursomanno, S., Aleliunaite, A., Wu, W., Mankouri, H.W., Shen, H., Liu, Y. and Hickson, I.D. (2015) Replication stress activates DNA repair synthesis in mitosis. Nature 528, 286-290.
- Nielsen, C.F., Huttner, D., Bizard, A.H., Hirano, S., Li, T-N., Palmai-Pallag, T., Bjerregaard, V.A., Liu, Y., Nigg, E.A., Wang, L.H-C. and Hickson, I.D. (2015) PICH promotes sister chromatid disjunction in mitosis: evidence of functional co-operation with topoisomerase II. Nature Communications 6, 8962..
- Ying, S., Minocherhomji, S., Chan, K.L., Palmai-Pallag, T., Chu, W.K., Wass, T., Mankouri H.W., Liu, Y. and Hickson, I.D. (2013) MUS81 promotes common fragile site expression. Nature Cell Biology 15, 1001-1007.
- Biebricher, A., Hirano, S., Enzlin, J.H., Wiechens, N., Streicher, W.W., Huttner, D., Wang, L.H-H., Nigg, E.A., Owen-Hughes, T., Liu, Y., Peterman, E., Wuite, G.J.L. and Hickson, I.D. (2013) PICH: A DNA translocase Specially Adapted for Processing Anaphase Bridge DNA. Molecular Cell 51, 691-701.
- Chan, K-L., Palmai-Pallag, T., Ying, S. and Hickson, I.D. (2009) Replication stress induces sister-chromatid bridging at fragile site loci in mitosis. Nature Cell Biology. 11, 753-760.
- Chu, W-K. and Hickson, I.D. (2009) RecQ helicases: multifunctional genome caretakers. Nature Reviews Cancer 9, 644-654
- Niels Mailand, Center for Protein Research, University of Copenhagen, Denmark.
- Dr. Keith Caldecott, Genome Damage and Stability Centre, University of Sussex, UK.
- Dr. Chuna Choudhary, Center for Protein Research, University of Copenhagen, Denmark.
- Dr Michael Lisby, University of Copenhagen, Denmark.
- Drs Erwin Peterman and Gijs Wuite, Free University of Amsterdam, The Netherlands.
- Dr Alessandro Vindigni, St. Louis University, USA.
- Dr Stephen West, Cancer Research UK, London Research Institute, UK
- Drs Songmin Ying and Huahao Shen, Zhejiang University, China
- Thanos Halazonetis, University of Geneva, Switzerland.
- Job Dekker, University of Massachusetts, USA.
- Bill Earnshaw, University of Edinburgh, UK.
- Masato Kanemaki, National Institute of Genetics, Shizuoka, Japan.
 Group Leader
Group Leader
Ian D. Hickson
Professor, PhD, FRS
Head of department
iandh@sund.ku.dk
(+45) 51 29 78 77
CV, publications, etc.
