Haahr group
Our lab aims to identify and characterize new genes that play important roles in human disease, such as cancer, by studying basic mechanisms of cellular stress responses and mRNA translation using a combination of unbiased proteome- and genome-wide screens as well as targeted molecular and cell biological approaches.

The molecular mechanisms of cellular stress responses
Cellular stress responses are comprised of dynamic signaling networks controlling specialized adaptation mechanisms that serve as a safeguard against various endogenous and environmental challenges, thereby promoting overall cell and organismal fitness. The precise regulation and proper execution of stress responses determines cell fate and are important for mitigating a myriad of severe malignancies, such as cancer and age-related neurodegenerative disorders. Particularly, we are interested in the tight interplay of these responses with the transcriptional and translational machineries that determine the gene expression output of the cell. One of our primary objectives entail the identification and characterization of new factors and mechanisms that underpin these responses at the molecular level, further elucidating how these systems are co-opted and disrupted in the context of malignancy, and ultimately devising strategies to manipulate them for therapeutic purposes.
We are addressing this challenge by combining targeted cell biology-, biochemistry-, and synthetic biology-driven approaches with cutting-edge systems-wide screening strategies to elucidate these processes in human cells.
Functional genomics
Essentially all human traits and diseases are shaped by variation in our genetic material. An integral part of our lab aims to implement and develop genetic strategies to couple genotype-phenotype relationships at scale to address our biological questions in new and better ways. Currently, we use a powerful combination of CRISPR-mediated genome engineering, ultra-deep genome mutagenesis, and FACS-based phenotypic screening to link genes to quantitative molecular traits in a high-throughput manner. A long term-goal of the lab is to further develop assays and multiplexed screening approaches to understand how genetic variation at the single nucleotide level regulates gene function and disease.
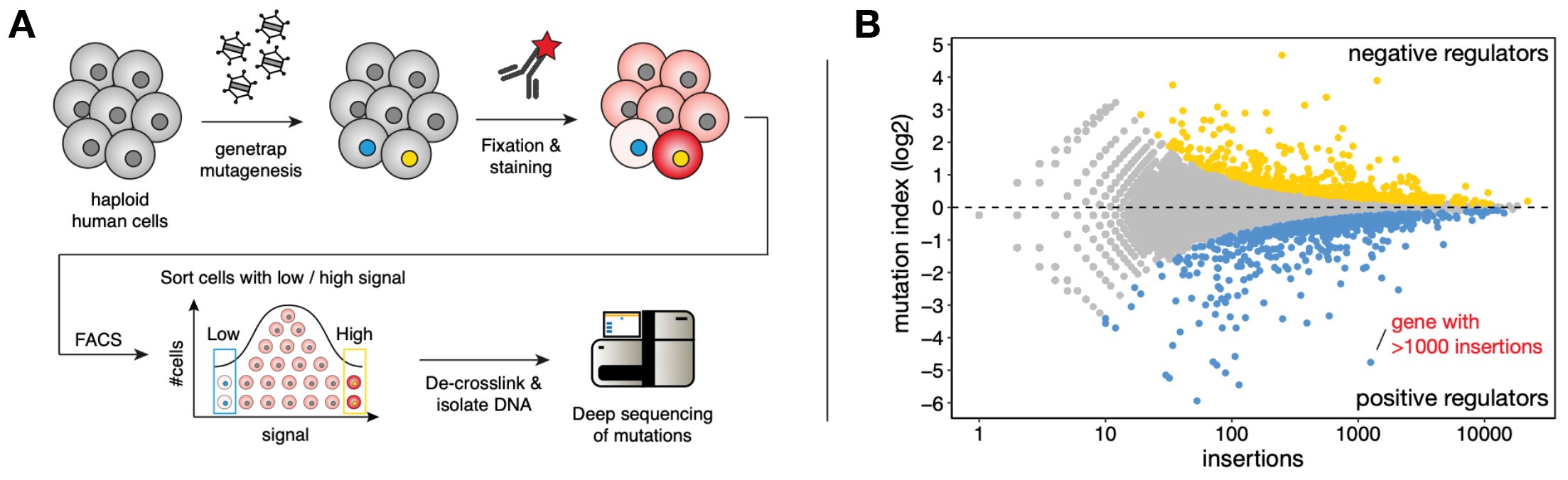
Pooled loss-of-function phenotypic screens in haploid human cells (A) Haploid human cells are mutagenized using a gene-trap retrovirus and subsequently fixed, permeabilized and stained using an antibody of interest. Cells are sorted using FACS to isolate distinct phenotypic populations (e.g. highest and lowest level of antibody staining). The isolated populations DNA is extracted, and mutations are identified by deep sequencing and assigned to genes. (B) Example of a screen fishtail plot displaying genes that cause increased antibody signal upon mutation (yellow dots, ‘negative regulators’) or lower amounts of antibody staining (blue dots, ‘positive regulators’).
- New mechanisms of mRNA translation regulation in cellular stress responses
- SUMO-targeted ubiquitination in proteostatic stress responses
- The cellular response to RNA damage and RNA-protein crosslinks in cancer
-
Haahr P, Galli RA, van den Hengel LG, Bleijerveld OB, Kazokaitė-Adomaitienė J, Song JY, Kroese LJ, Krimpenfort P, Baltissen MP, Vermeulen M, Ottenheijm CAC, Brummelkamp TR. Actin maturation requires the ACTMAP/C19orf54 protease. Science. 2022 Sep 30;377(6614):1533-1537.
-
Nordgaard C, Vind AC, Stonadge A, Kjøbsted R, Snieckute G, Antas P, Blasius M, Reinert MS, Del Val AM, Bekker-Jensen DB, Haahr P, Miroshnikova YA, Mazouzi A, Falk S, Perrier-Groult E, Tiedje C, Li X, Jakobsen JR, Jørgensen NO, Wojtaszewski JF, Mallein-Gerin F, Andersen JL, Pennisi CP, Clemmensen C, Kassem M, Jafari A, Brummelkamp T, Li VS, Wickström SA, Olsen JV, Blanco G, Bekker-Jensen S. ZAKβ is activated by cellular compression and mediates contraction-induced MAP kinase signaling in skeletal muscle. EMBO J. 2022 Sep 1;41(17):e111650.
-
Liu JCY, Kühbacher U, Larsen NB, Borgermann N, Garvanska DH, Hendriks IA, Ackermann L, Haahr P, Gallina I, Guérillon C, Branigan E, Hay RT, Azuma Y, Nielsen ML, Duxin JP, Mailand N. Mechanism and function of DNA replication-independent DNA-protein crosslink repair via the SUMO-RNF4 pathway. EMBO J. 2021 Sep 15;40(18):e107413
-
Achuthankutty D*, Thakur RS*, Haahr P*, Hoffmann S, Drainas AP, Bizard AH, Weischenfeldt J, Hickson ID, Mailand N. Regulation of ETAA1-mediated ATR activation couples DNA replication fidelity and genome stability. J Cell Biol. 2019 Dec 2;218(12):3943-3953. *shared first author
-
Haahr P, Borgermann N, Guo X, Typas D, Achuthankutty D, Hoffmann S, Shearer R, Sixma TK, Mailand N. ZUFSP Deubiquitylates K63-Linked Polyubiquitin Chains to Promote Genome Stability. Mol Cell. 2018 Apr 5;70(1):165-174.e6. doi: 10.1016/j.molcel.2018.02.024.
-
Haahr P, Hoffmann S, Tollenaere MA, Ho T, Toledo LI, Mann M, Bekker-Jensen S, Räschle M, Mailand N. Activation of the ATR kinase by the RPA-binding protein ETAA1. Nat Cell Biol. 2016 Nov;18(11):1196-1207.
 Group Leader
Group Leader
Peter Thorlund Haahr
associate professor
pha@sund.ku.dk
(+45) 35 33 06 80
CV, Publications, etc.
Transcription, RNA, and Gene Medicine Program
Center for Gene Expression
