Halim Group
Our group is interested in the biosynthesis, regulation and biochemistry of O-linked glycosylations with a special focus on protein O-mannosylations.
Glycosylation is an essential cellular process that involves the covalent attachment of glycans (carbohydrates/sugars) to proteins. Glycans play important roles in maintaining normal cellular functions and dysregulation of processes related to protein glycosylations are known to cause various diseases, e.g. cancer and developmental disorders. Our group is interested in the biosynthesis, regulation and functions of O-linked glycosylations with a special focus on protein O-mannosylation (O-Man). We use a combination of methods, including advanced mass spectrometry, to study structures, map site-specific locations and quantify changes of protein O-mannosylations on a proteome-wide scale. In addition, we use CRISPR/Cas9 gene editing for knock-out/knock-in of glycogenes for dissection of enzymatic pathways and characterization of disease variants in cell-based models.

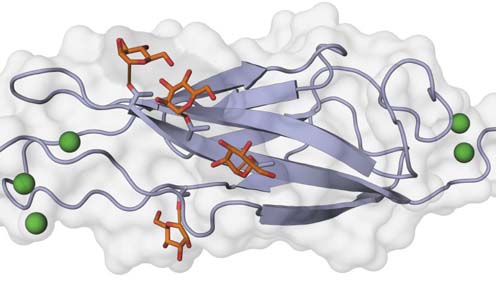
Cadherin O-mannosylation – the fourth extracellular cadherin (EC) domain of mouse E-cadherin is shown with O-linked mannose glycans (sticks) on specific β-strands. Calcium ions are depicted as green spheres. Illustration by Oliver Harrison, Julia Brasch and Lawrence Shapiro (Columbia University).
Cadherin-specific O-mannosylation by TMTC1-4
O-mannosylation was for a long time believed to be a rare type of protein modification in mammals. Until recently, α-dystroglycan (αDG), a component of the dystrophin complex, remained the only well-characterized protein with respect to O-mannosylation sites and structures. We developed a glycoproteomic workflow for studying protein O-mannosylations on a proteome-wide scale, which allowed us to greatly expand the human O-mannose glycoproteome and led to the discovery of cadherins and protocadherins as major carriers of O-mannosylations. Furthermore, we uncovered that O-mannosylation of the cadherin superfamily is not mediated by the classical POMT1/POMT2 enzymes and identified the glycosyltransferase family (GT105), composed of TMTC1-4, which initiates the cadherin-specific O-mannosylation.
Currently, we are characterizing the TMTC1-4 enzyme family, exploring the substrate specificities of individual TMTC family members and studying the disease-causing mutations in this enzyme family. Our ambitions are to gain further understanding on cellular processes related to the function of cadherin-specific O-mannosylation in health and disease (e.g. Cobblestone lissencephaly).
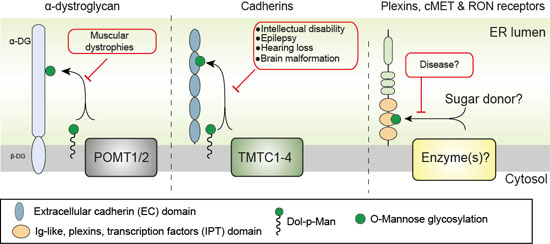
Differential regulation of protein O-mannosylation in metazoans. O-mannosylation is predicted to be controlled by at least three distinct enzyme families; α-dystroglycan O-mannosylation is initiated by the POMT1/POMT2 family, the cadherin superfamily is glycosylated by the TMTC1-4 family and the plexin family (IPT domains) is O-mannosylated by unknown enzyme(s). Modified from Larsen et al. (2019) Curr Opin Struct Biol
Identification of new genes and pathways involved in O-mannosylation and cell-cell communication
In addition to the cadherin superfamily, we have identified plexin receptors as a major class of cell-surface O-mannosylated proteins. The IPT domains of plexins are frequently O-mannosylated on specific β-strands but, intriguingly, this O-mannosylation doesn’t seem to be mediated by POMT1/POMT2 or the TMTC1-4 family. Our results thus indicate that other, as yet unknown glycosyltransferases are present in mammalian systems and are responsible for directing O-mannosylation specifically to proteins with IPT folds. We are currently using bioinformatics tools for prediction and screening of candidate genes to test this hypothesis and to identify the novel glycosyltransferase enzymes responsible for O-mannosylation in mammalian cells.
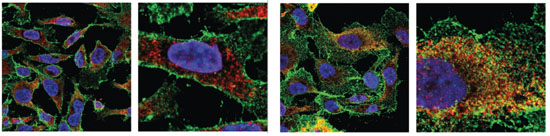
O-Man glycoproteome discovery
The evolution of multiple O-glycosylation pathways was likely driven by a need to adapt higher organism to their environment and for their cells/proteins to obtain more specialized properties and functions, tunable via different types of O-glycans, to control e.g. protein trafficking, cell adhesion, protease cleavage and signaling. However, the precise roles of O-Man glycans on specific proteins are largely unknown, mainly due to a lack in the understanding of which proteins and sites are modified by O-Man glycans (the O-Man glycoproteome) and how this specificity is achieved.
Developing new techniques capable of decoding the regulations and functions of O-Man glycoproteomes and may thus lead to the next breakthrough in the understanding of how cellular functions are regulated and fine-tuned by O-glycans, how O-Man glycoproteomes contribute to health and disease, and ultimately, how this important class of molecules could be exploited for development of new strategies for diagnostics, drugs and treatments.
In this project, we develop sensitive glycopeptide enrichment protocols for site-specific, quantitative O-Man glycoproteomics in cell models and tissues. Specific applications include O-Man discovery in model organisms, e.g. Drosophila melanogaster and mice (brain).
O-Man dependent interactomics
Adhesion molecules and receptors engage in complex physical interactions that rely on extensive protein-protein interaction networks. However, insight into how the novel O-Man glycans on distinct protein domains influence cadherin and plexin receptor interaction is completely lacking. Testing specific hypotheses for glycoproteins in general has remained challenging, and while great progress has been made towards understanding direct glycan-protein binding, far less is known about how glycosylation status diversifies protein-protein interactions (interactomes).
We aim to develop novel analytical platforms integrating precise gene editing, endogenous tagging, glycoproteomics and interactomics to address an important unmet need for new, innovative solutions capable of resolving glycoprotein interactomes in the context of their (differential) glycosylation status.
The mucin-like KIAA1549 protein
Genetic defects involving the KIAA1549 gene underlie human disorders, including Pilocytic Astrocytoma (PA), which accounts for the most common form of pediatric brain tumors. The most common genetic alternation in PA results in a KIAA1549-BRAF kinase fusion protein with constitutive BRAF kinase activity which impacts the MAPK/RAS signaling pathway. However, the molecular functions of KIAA1549 remain unknown, and this knowledge gap hampers further advances in the understanding of molecular mechanisms in pediatric cancers.
We previously discovered that KIAA1549 is a heavily O-Man glycosylated protein and a target for the POMT1/POMT2 pathway. The overarching objective in this project is to apply genetic engineering, mass spectrometry and cell models to characterize the O-Man glycosylation profile and molecular functions of KIAA1549 in health and disease.
- Povolo L, Tian W, Vakhrushev SY, Halim A. (2024) Global View of Domain-Specific O-Linked Mannose Glycosylation in Glycoengineered Cells. Mol Cell Proteomics. 2024 Jul;23(7):100796.
- Larsen ISB, Povolo L, Zhou L, Tian W, Mygind KJ, Hintze J, Jiang C, Hartill V, Prescott K, Johnson CA, Mullegama SV, McConkie-Rosell A, McDonald M, Hansen L, Vakhrushev SY, Schjoldager KT, Clausen H, Worzfeld T, Joshi HJ, Halim A. (2023) The SHDRA syndrome-associated gene TMEM260 encodes a protein-specific O-mannosyltransferase. PNAS 2023 May 23;120(21):e2302584120.
- Ieva Bagdonaite, Stacy A. Malaker, Daniel A. Polasky, Nicholas M. Riley, Katrine Schjoldager, Sergey Y. Vakhrushev, Adnan Halim, Kiyoko F. Aoki-Kinoshita, Alexey I. Nesvizhskii, Carolyn R. Bertozzi, Hans H. Wandall, Benjamin L. Parker, Morten Thaysen-Andersen & Nichollas E. Scott (2022) Glycoproteomics Nature Reviews Methods Primers volume 2, Article number: 48 (2022)
- Larsen ISB, Narimatsu Y, Clausen H, Joshi HJ, Halim A. (2019) Multiple distinct O-Mannosylation pathways in eukaryotes. Curr Opin Struct Biol. 2019 Jun;56:171-178.
- Larsen, I. S. B., Narimatsu, Y., Joshi, H. J., Siukstaite, L., Harrison, O. J., Brasch, J., Goodman, K. ., Hansen, L., Shapiro, L., Honig, B., Vakhrushev, S. Y., Clausen, H., and Halim, A. (2017) Discovery of an O-mannosylation pathway selectively serving cadherins and protocadherins. PNAS 114, 11163-11168
- Larsen, I. S. B., Narimatsu, Y., Joshi, H. J., Yang, Z., Harrison, O. J., Brasch, J., Shapiro, L., Honig, B., Vakhrushev, S. Y., Clausen, H., and Halim, A. (2017) Mammalian O-mannosylation of cadherins and plexins is independent of protein O-mannosyltransferases 1 and 2. J Biol Chem 292, 11586-11598
- Rubinstein R, Thu CA, Goodman KM, Wolcott HN, Bahna F, Mannepalli S, Ahlsen G, Chevee M, Halim A, Clausen H, Maniatis T, Shapiro L, Honig B. (2015) Molecular logic of neuronal self-recognition through protocadherin domain interactions. Cell. 2015 Oct 22;163(3):629-42
- Vester-Christensen, M. B., Halim, A., Joshi, H. J., Steentoft, C., Bennett, E. P., Levery, S. B., Vakhrushev, S. Y., and Clausen, H. (2013) Mining the O-mannose glycoproteome reveals cadherins as major O-mannosylated glycoproteins. PNAS 110, 21018-21023
- John LaCava (The Rockefeller University, NY, USA)
- Thomas Worzfeld (University of Marburg, Germany)
- Vladislav Panin (Texas A&M University, USA)
- Joseph Gleeson (University of California San Diego, USA)
We welcome applications from motivated postdoctoral and predoctoral researchers as well as master and bachelor students. Please contact us at halim@sund.ku.dk
Techniques and training platforms:
- Genetic engineering (knock-out/knock-in) by CRISPR/Cas9 and Zinc Finger Nucleases
- CRISPR/Cas9-based endogenous tagging of mammalian proteins
- Protein expression in bacteria and mammalian cells
- Protein purification and glycopeptide enrichment (HPLC, AKTA, size exclusion)
- Mass spectrometry (MALDI-TOF, Orbitrap Fusion/Lumos)
- Omics analyses (Proteomics, Glycomics, Glycoproteomics)
- Functional characterization of protein glycosylation
- Cell-based disease model







 Group Leader
Group Leader