Tommerup Group
The research focus of Tommerup Group is to use balanced chromosomal rearrangements (BCR) to identify and characterize novel disease and phenotype genes, regulatory domains (Topological Associating Domains (TADs)) of disease genes and novel genetic mechanisms, including X-inactivation and germline chromothripsis. Sex differential responses to Sars-Cov-2-infections. Meiosis in Nature, where we by a new spreading technique can visualize meiotic chromosomes in wild animals, and establish their karyotype as a supplement to genome sequencing.
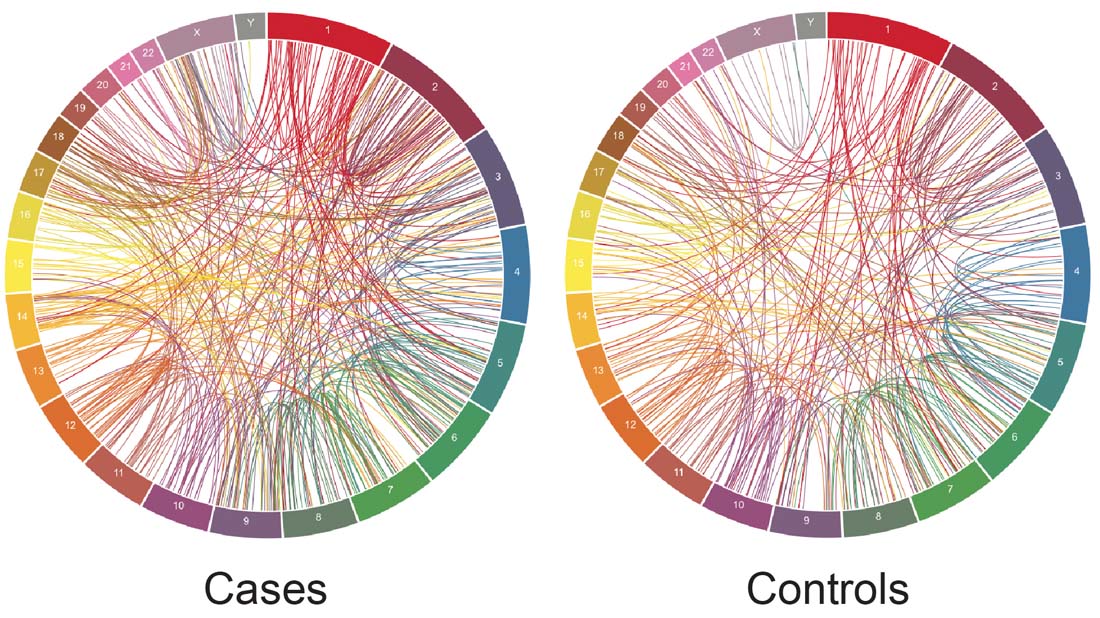
Circa plots of the genomic distribution of breakpoints in >700 sequence resolved simple two-way balanced chromosomal rearrangements.
We study balanced chromosomal rearrangements (BCR) to identify and characterize novel disease genes. As part of this we look at genetic mechanism related to BCRs, including 1) long-range position effects due to truncation of regulatory genomic domains, e.g Topological Associating Domains (TADs) and chromatin loops; 2) X-inactivation; and 3) germline chromothripsis. Recently, we initiated “Meiosis in Nature”, to study meiosis in wild animals as a way to complement The Earth Biogenome Project, the humongous effort to sequence all living species. Two examples illustrate the potential: 1) meiotic karyotyping of the Greenland shark (Somniferus microcephalus), the longest living vertebrate known; 2) meiosis in sika (Cervus nippon) and red deer (Cervus elapus) and in potential hybrids between them.
- The first study of long-term risk of prenatally diagnosed de novo BCRs increases the risk from 7% to appr. 20% and lead to new guidelines (Am J Hum Genet 2018;102:1090-1103)
- The multiple breakpoints in germline chromothripsis may predispose to complex multigenic disorders (Hum Mutat 2019;40:1057-62)
- Haploinsufficiency of ARHGAP42 leads to age-dependent hypertension (Eur J Hum Genet 2019;27:1296-1303). The study also provides support for an obesity locus in the CELF4 regulatory domain on chromosome 18
- Combining DNA-DNA-interaction (Hi-C) studies with short and long read sequencing improve the dissection of germline complex chromosomal rearrangements (Nat Commun. 2022;13:6470)
- Maternal smoking delays fetal X-inactivation (Reprod Toxicol. 2025;137:109010)
- Multicenter study pin-point glycosylation processes in human genetic susceptibility to infections with SARS-CoV-2 Omicron variants (Nat Genet 2026;58:299–306)
- Coordination of International Breakpoint Mapping Consortium (2014-), involving multiple diagnostic cytogenetic laboratories from >50 countries/6 continents.
- We have accumulated the largest number (~800) of sequence resolved germline BCRs, including the first large control BCR cohort (collaboration with Michael Talkowksi (Harvard) and Chelsea Lowther (Mount Sinai, New York). Direct gene truncation may explain ~18% of BCR-associated early developmental disorders (DD). We provide the first ranked list of TADs as genomic units for BCR-mediated long-range-position effects (LRPE). We define novel LRPE-loci (e.g. BCL11A, BCL11B) and our data suggest that LRPE may be a frequent cause of DD in BCRs. This window into the developmental regulome establish links to multiple disorders, incl. intellectual disability, autism, epilepsy, narcolepsy, speech defects and congenital brain malformations (in revision for Nature Genet).
- We have initiated the first systematic X-inactivation study of sequence resolved X;autosomal translocations and X-inversions.
- We are studying the largest cohort of cases with complex germline chromothripsis (collaboration with guest researcher Lusine Nazaryan-Petersen and Anna Lindstrand, Karolinska, Sweden).
- Search for host genetic factors that may be underlie sex differential responses to Sars-Cov-2, with a particular focus on the X-chromosome.
- In “Meiosis in Nature”, we study meiotic chromosomes in wild animals as a way to complement The Earth Biogenome Project, the ongoing effort to sequence all living species. Our first selected organism is the Greenland shark ✝, the longest living vertebrate known.
✝John Fleng Steffensen in memoriam
345 published peer reviewed publications, >30,000 citations, h-index: 81
Google Scholar
- The Danish Council for Independent Research - Medical Sciences [4183-00482B]. International Breakpoint mapping Consortium.
- The Danish Council for Independent Research - Medical Sciences [0216-00014B). COVID-19 sex difference: X-linked genetic variants affecting disease trajectories and survival.
- Horizon Europe. Project 101057129. Respiratory Host-Pathogen Interaction (REACT).
 Group Leader
Group Leader
Niels Tommerup
Senior Researcher, DMSc
ntommerup@sund.ku.dk
(+45) 30 31 43 86
CV, publications, etc.
